The First Multi-Institutional Analytical Demonstration of a Spatial Biology Workflow published in the Journal for ImmunoTherapy of Cancer
Synopsis: The PhenoImager™ (formerly called Phenoptics) mIF solution was used in a multi-site study to demonstrate and validate an automated end-to-end workflow that characterizes PD-1/PD-L1 immune checkpoint signaling in tumor tissue samples. The paper titled, “Multi-institutional TSA-amplified Multiplexed Immunofluorescence Reproducibility Evaluation (MITRE Study),” was published in the Journal for ImmunoTherapy of Cancer (JITC) in July 2021. The MITRE results are an important step toward standardizing an automated mIF-based spatial biology workflow that provides the level of performance needed to support clinical trials and that can be applied to clinical testing in the future.
Top immuno-oncology and pathology experts from five institutions collaborated with Akoya to conduct the MITRE study, including Johns Hopkins University School of Medicine, Yale University School of Medicine, Earle A. Chiles Research Institute, The University of Texas MD Anderson Cancer Center, and Bristol Myers Squibb.
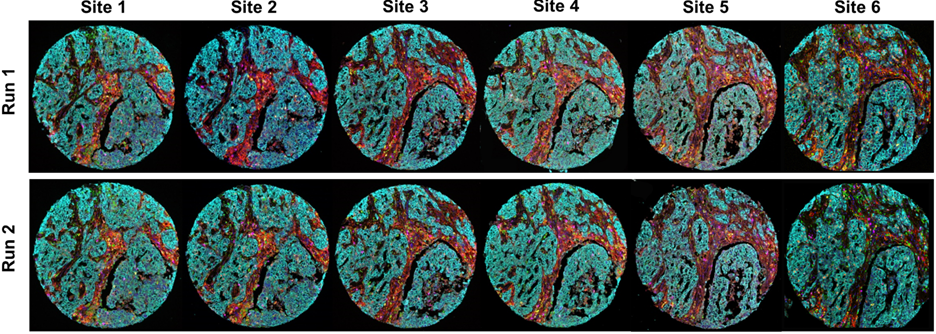
Shown here are multiplex immunofluorescence images, from all the sites involved in the study, where six biomarkers were analyzed in breast cancer samples.
Spatial biomarkers outperform PD-L1 IHC and TMB in predicting immunotherapy response
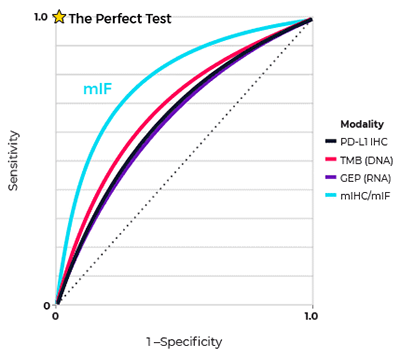 Synopsis: In a seminal multi-institutional study, published in JAMA Oncology in 2019 , a group of leading immuno-oncology experts determined that spatial phenotypic signatures, measured by multiplex immunofluorescence (mIF), outperformed other biomarker testing approaches in predicting response to anti-PD-1/PD-L1 treatments. The study was conducted by scientists at Johns Hopkins University, Yale University, Vanderbilt University, and Northwestern University.
Synopsis: In a seminal multi-institutional study, published in JAMA Oncology in 2019 , a group of leading immuno-oncology experts determined that spatial phenotypic signatures, measured by multiplex immunofluorescence (mIF), outperformed other biomarker testing approaches in predicting response to anti-PD-1/PD-L1 treatments. The study was conducted by scientists at Johns Hopkins University, Yale University, Vanderbilt University, and Northwestern University.
The authors reviewed published data from more than 50 studies covering more than 10 types of cancer and over 8,000 patients. The mIF studies featured in this paper cite the use of Akoya’s PhenoImager™ (formerly called Phenoptics) platform.
Astronomy Meets Pathology: A novel spatial phenotypic signature to predict immunotherapy response and outcomes in melanoma
 Synopsis: In a truly unique approach taken by scientists at Johns Hopkins University (JHU), Dr. Janis Taube, a leading pathology expert and Dr. Alex Szalay, a world-renowned astrophysicist, joined forces to solve the big data challenge in cancer biomarker discovery. They built a platform titled, AstroPath™ and applied celestial object mapping algorithms to the study of the tumor microenvironment, enabling rapid probabilistic studies of how tumor and immune cells organize and interact to influence treatment response.
Synopsis: In a truly unique approach taken by scientists at Johns Hopkins University (JHU), Dr. Janis Taube, a leading pathology expert and Dr. Alex Szalay, a world-renowned astrophysicist, joined forces to solve the big data challenge in cancer biomarker discovery. They built a platform titled, AstroPath™ and applied celestial object mapping algorithms to the study of the tumor microenvironment, enabling rapid probabilistic studies of how tumor and immune cells organize and interact to influence treatment response.
The underlying image-capture technology for AstroPath is the PhenoImager™ (formerly called Phenoptics) platform, as part of an ongoing collaboration between Akoya and JHU. The result was the groundbreaking discovery of a spatial biomarker signature which is highly predictive of immunotherapy response in melanoma cases. The study was published in Science in 2021.
A seminal concept in spatial analysis of tissue samples: Cellular Neighborhoods.
 Synopsis: Dr. Garry Nolan of Stanford University, a well-recognized thought leader in immunology and spatial biology, published a novel analytical framework to uncover “cellular neighborhoods” in the tumor microenvironment. In simple terms, “the immune tumor microenvironment is like a city of neighborhoods (e.g., industrial, residential, or agricultural), which are regions where specific functions of the city occur.” The Stanford team demonstrated how the organization of these neighborhoods was predictive of patient outcomes in colorectal cancer, underscoring the importance of spatial context in developing prognostic biomarkers.
Synopsis: Dr. Garry Nolan of Stanford University, a well-recognized thought leader in immunology and spatial biology, published a novel analytical framework to uncover “cellular neighborhoods” in the tumor microenvironment. In simple terms, “the immune tumor microenvironment is like a city of neighborhoods (e.g., industrial, residential, or agricultural), which are regions where specific functions of the city occur.” The Stanford team demonstrated how the organization of these neighborhoods was predictive of patient outcomes in colorectal cancer, underscoring the importance of spatial context in developing prognostic biomarkers.
This seminal spatial biology algorithm was developed with data generated solely on Akoya’s PhenoCycler™ (formerly called CODEX) system and the results were published in Cell in 2020. The analytical framework presented in this paper serves as an industry-standard for how researchers can derive actionable insights from the billions of data points generated from each tissue image.
Studying the biology of tumor-immune interactions in colorectal cancer
Speaker: Jérôme Galon, PhD, INSERM, Paris
Synopsis: ImmunoScore is one of the first spatial biomarker signatures listed in the WHO and ESMO guidelines and is used to refine the prognosis of patients with localized colon cancer (Stage II and III cases). Dr. Galon, Director of the Laboratory of Integrative Cancer Immunology at INSERM, Paris, pioneered the development of the ImmunoScore test.
In this Science webcast, Dr. Galon gives an educational overview of the immune landscape of cancer and provides a summary of latest studies to profile ImmunoScore and other immune signatures in metastatic colorectal cancer (Stage IV cases). Leveraging a suite of multi-omics tools, Dr. Galon cites the use of Akoya’s multispectral imaging technology, PhenoImager™ (formerly called Phenoptics), to quantify the spatial relationships of diverse cell phenotypes in the tumor microenvironment.
Studying the biology of tumor-immune interactions in breast cancer samples from the I-SPY 2 trial
Speaker: Alexander Borowsky, MD, UC Davis
Synopsis: Akoya, UCSF and the I-SPY trial consortium have an ongoing collaboration to leverage the PhenoImager™ (formerly called Phenoptics) platform for developing predictive and prognostic biomarkers, particularly for use in selecting the most effective neoadjuvant and adjuvant immunotherapies for patients with early breast cancer. At the Society for Immunotherapy of Cancer 2020 Annual Meeting, Dr. Borowsky presented preliminary results from the I-SPY2 trial where he showed how spatial proximity scores between tumor and immune cells are correlated to treatment response.
Adding spatial context to the Human Cell Atlas Initiative: Breast Atlas Case Study
Speaker: Kai Kessenbrock, PhD, UC Irvine
Synopsis: The mission of the Human Cell Atlas is to create comprehensive reference maps of all human cells to describe and define the cellular basis of health and disease. Using the Human Breast Cell Atlas project as an example, Dr. Kessenbrock explains how his team discovered unique cellular niches within the breast tissue microenvironment and how their spatial proximity gives us a more comprehensive view of the biology underlying each sample.
Dr. Kessenbrock’s lab complements single-cell RNA sequencing data with single-cell spatial mapping data from the PhenoCycler™ (formerly called CODEX) platform to generate an atlas of human breast tissue.
Spatial Multi-Omics Webinar Series– Complementing single-cell RNA-Sequencing data with Spatial Imaging data
Synopsis: In this multi-part webinar series, the speakers review analytical frameworks and algorithms to integrate imaging-based single-cell data with complementary transcriptomic and genomic datasets.
The experts combined PhenoCycler™ (formerly called CODEX) with single-cell sequencing data to generate multi-omic spatial maps of tissue samples. The speakers cite the ability of the PhenoCycler™ (formerly called CODEX) platform to generate high resolution maps of millions of cells across whole tissue samples as one of the key reasons for integrating it with genomic datasets.
Cancer Immunome Project – Adding the Spatial Dimension
 Interviewee: JC Villasboas, MD, Mayo Clinic
Interviewee: JC Villasboas, MD, Mayo Clinic
Synopsis: Dr. JC Villasboas, a physician-scientist and Director of the Immune Monitoring Core Facility at the Mayo Clinic, and his team of collaborators are developing what they call The Cancer Immunome Project. This is a comprehensive effort to fully characterize the immune system and how it interacts with and fights off cancer.
In the interview, Dr. Villasboas says the immune system is of such complexity that it took the addition of new spatial biology tools, namely Akoya’s PhenoCycler™ (formerly called CODEX) system—to be able to comprehensively tackle such a project.






